The OpenNLP Custom NER Model Extraction Engine
This engine allows the configuration of custom Apache OpenNLP NameFinder models for NER of plain text content.
Example Result
This engine adds fise:TextAnnotation for the processed plain text to the metadata of the content item. The following code listing shows an DNA type Named Entity detected based on a OpenNLP NameFinder model trained based on the BioNLP2004 dataset:
{ "@subject": "urn:enhancement-0e31eb01-23c5-82b5-1372-5c5606c09960", "@type": [ "Enhancement", "TextAnnotation" ], "confidence": 0.40148407, "creator": "org.apache.stanbol.enhancer.engines.opennlp.impl.CustomNERModelEnhancementEngine", "start": 228, "end": 242, "extracted-from": "urn:content-item-sha1-84a30aeeb073be543f7c54266e232aae572efac0", "selected-text": { "@language": "en", "@literal": "HIV-2 enhancer" }, "selection-context": { "@language": "en", "@literal": "activation of the HIV-2 enhancer in monocytes and T cells" }, "type": "http://www.bootstrep.eu/ontology/GRO#DNA" },
Configuration
The usage of this Engine requires to create a service configuration. Configurations require at least a single NameFinderModel name to be configured.
Parameters
- Name Finder Models (stanbol.engines.opennlp-ner.nameFinderModels): The list if custom NameFinderModels used by this engine. The Engine supports Arrays, Vectors and comma separated string for. Values are the file names of the NameFinderModel files. Configured files are loaded by using the DataFileProvider service. That means that files copied into the 'datafile' folder (by default located at '{stanbol-working-dir}/stanbol/datafiles').
- Named Entity to 'dc:type' Mappings (stanbol.engines.opennlp-ner.typeMappings): This configuration uses the syntax {named-entity-type} > {uri}": {named-entity-type} matches to the string "name" used for the named entity type in the OpenNLP NameFinder model. {uri} MUST BE a valid URI and is used as dc:type value for fise:TextAnnotations created by the engine for extracted Named Entities. NOTE: that TextAnnotations for unmapped Named Entity Types will have no dc:type information.
The following figure provides a visual representation of an engine configuration configured for all NamedEntity types supported by the BioNLP2004 dataset.
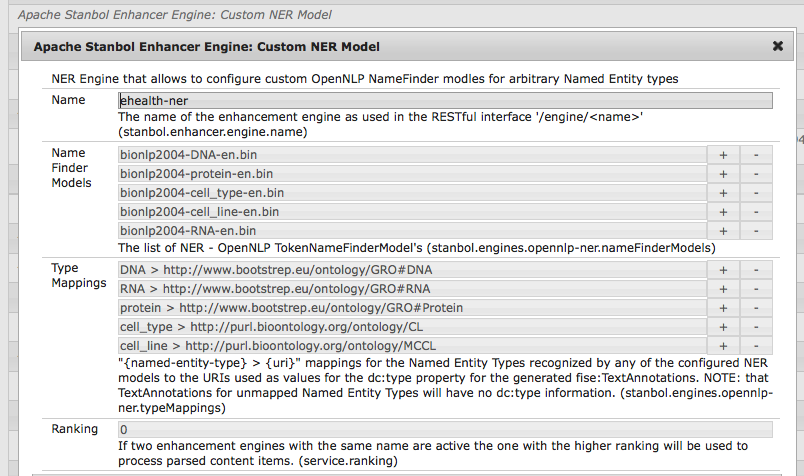
The same configuration can be also provided as OSGI configuration file with the name 'org.apache.stanbol.enhancer.engines.opennlp.impl.CustomNERModelEnhancementEngine-ehealthner.config' and the contents:
stanbol.enhancer.engine.name="ehealth-ner" stanbol.engines.opennlp-ner.nameFinderModels=["bionlp2004-DNA-en.bin","bionlp2004-protein-en.bin","bionlp2004-cell_type-en.bin","bionlp2004-cell_line-en.bin","bionlp2004-RNA-en.bin"] stanbol.engines.opennlp-ner.typeMappings=["DNA\ >\ http://www.bootstrep.eu/ontology/GRO#DNA","RNA\ >\ http://www.bootstrep.eu/ontology/GRO#RNA","protein\ >\ http://www.bootstrep.eu/ontology/GRO#Protein","cell_type\ >\ http://purl.bioontology.org/ontology/CL","cell_line\ >\ http://purl.bioontology.org/ontology/MCCL"]
NOTE: that the '.config' format requires spaces to be escaped with '\'